In the field of oceanography, statistical analysis of complex data is a fundamental skill that often calls for mathematics beyond the training of the student. Here I would like to walk through some simple linear algebra and connect it to a well-used yet poorly understood statistical analysis technique.
Eigenpairs
Given a linear transformation (e.g. $x\mapsto x’$), there exists a matrix which does this mapping: $Ax = x’$. To illustrate, let’s use the example: $$x= \begin{pmatrix} 1 \\ 2 \end{pmatrix} \text{ and } A = \begin{pmatrix} 2 & 1 \\ 1 & 2 \end{pmatrix}$$
This transformation (i.e. $A$) maps the vector $x$ to: $$x’=\begin{pmatrix} 4 \\ 5\end{pmatrix}$$

With this example transformation, $A$, let’s take a look at eigenvalues and eigenvectors.
An eigenvector is a vector whose direction is preserved by the transformation. In other words, the vector is only scaled by the transformation and still points in the same direction (or 180º off). In the example above, we can see that the black vector its transformation in red do not point in the same direction, therefore the black vector is not an eigenvector of $A$.
To find the eigenvectors, we set up a simple equation that summarizes our definition above. The equation below simply states that the transformation of $\vec{x}$ must be a multiple of $\vec{x}$ (i.e. same direction but scaled by $\lambda$).
$$A\vec{x} = \lambda \vec{x}$$
Rearranging the equation we find that $(A-I)\vec{x} = 0$. In order for this equation to hold, the determinant $|A-I|$ must also equal 0. While the details on developing the characteristic polynomial are not to be covered here[1], the solution of this equation yields values of $\lambda$ which are intimately connected to the eigenvector. We term these constants the eigenvalues of $A$ and represent the magnitude of the scaling between the eigenvector and its transformation (i.e. how the length changes).
Covariance Matrix
Starting with a set of data, such as student test scores, we can construct a data matrix that we will call $A$. For the example shown below, we’ve placed each student’s test scores in the data matrix so that student 1 received an 80 on the first test, 72 on the second test, and a 90 on the third test. Let’s call the first test (column 1) the Math test, the second an English test, and the last column the History test.
$$A = \begin{pmatrix} 80 & 72 & 90 \\ 74 & 67 & 68 \\ 90 & 88 & 93 \\ 67 & 88 & 75 \end{pmatrix}$$
Looking at the matrix here, there are 4 students and three test; but we could easily add more students (rows) or more tests (columns) without too much difficulty. The covariance matrix will be a 3×3 matrix relating how the score on test is correlated to the other tests. This will become clearer once we’ve finished.
To calculate the covariance matrix we simply follow the following formula:
$$C = \frac{1}{n-1}A’^TA’$$
where $A’ = A-\frac{1}{n}(1)A$. The first transformation from $A\to A’$ removes the average value from each of the columns (in technical terms it centers the data). For our data matrix, $A$, this becomes:
$$A’ = \begin{pmatrix} 80 & 72 & 90 \\ 74 & 67 & 68 \\ 90 & 88 & 93 \\ 67 & 88 & 75 \end{pmatrix} – \frac{1}{4} \begin{pmatrix} 1&1&1&1\\1&1&1&1\\1&1&1&1 \end{pmatrix} \begin{pmatrix} 80 & 72 & 90 \\ 74 & 67 & 68 \\ 90 & 88 & 93 \\ 67 & 88 & 75 \end{pmatrix} = \begin{pmatrix}2.25 & -6.75 & 8.5 \\ -3.75 & -12.25 & -13.5 \\ 12.25 & 9.25 & 11.5 \\ -10.75 & 9.25 & -6.5\end{pmatrix}$$
The next step is to left multiply $A’$ with its transpose:
$$C=\frac{1}{4-1}\begin{pmatrix}2.25 & -3.75 & 12.25 & -10.75 \\ -6.75 & -12.25 & 9.25 & 9.25 \\ 8.5 & -13.5 & 11.5 & -6.5\end{pmatrix}\begin{pmatrix}2.25 & -6.75 & 8.5 \\ -3.75 & -12.25 & -13.5 \\ 12.25 & 9.25 & 11.5 \\ -10.75 & 9.25 & -6.5\end{pmatrix} = \begin{pmatrix}1.00 & 0.47 & 0.75\\0.47 & 1.00 & 0.07\\0.75 & 0.07 & 1.00\end{pmatrix}$$
So now we have our covariance matrix, $C$; what does it actually tell us? Well, the covariance matrix will always be symmetric with a diagonal of zeros. It’s in the off-diagonal entries that tell us something. In this example we see that the third entry is 0.47, which says that the correlation between test 1 and test 3 is 0.75. Looking back at our data matrix, we can make sense of this number. Notice that a student who scored well in the first test often scored high on the third test, and thus a strong, positive correlation coefficient (0.75).
Looking at the correlation between the second and third test we see a coefficient of 0.07, which is very close to 0.0 meaning that those test scores were uncorrelated (doing well on one doesn’t say how well the student would do in the other).
Armed with our knowledge of eigenpairs and the covariance matrix, we can now look at the underpinning of the PCA analysis.
Putting it all together
Using the formula for eigenvalues outlined above, let’s make a table of the eigenpairs for our covariance matrix $C$.
$$\begin{matrix}\text{Entry} &\text{Value} & \text{Vector} \\1 & 1.92 & <-0.69, -0.40,-0.60>\\2 & 0.94 & <0.04, -0.85, 0.53 \\ 3 & 0.15 & <0.72, -0.34,-0.60>\end{matrix}$$
With these three eigenvectors and their corresponding eigenvalues, we can now run our PCA analysis. By definition eigenvectors are all orthogonal and therefore can be used to form a new basis for the data (hence an EOF method). In addition, the eigenvalue provides information on how representative that direction is according to the data. Here the first entry has the largest eigenvalue by far (1.92), and therefore accounts for the largest proportion of the variance within the data. To put it another way, if we wanted to summarize the trends in the data with the least amount of information, eigenvector 1 is your answer.
Eigenvector 1, $\vec{e_1}=<-0.69, -0.40,-0.60>$, says that if you lower one test score, it is a good bet that the other test scores will go down too (NB all entries in eigenvector are negative and of similar magnitude). The other eigenvectors provide other information on relations within the data, but due to the lower eigenvalues their contribution is less important.
The purpose of this analysis is to find a smart way to reduce the dimensionality of the data while preserving the maximum of information about the trends. The example data here is three dimensional, but if we wanted to plot it in two dimensions, then projecting it based on the first two eigenvectors (i.e. the ones with the largest eigenvalues) would preserve the greatest amount of information or variance.
When performing a PCA analysis, you’re generally working with high dimensional data so that instead of three eigenpairs, there might be a dozen or more: try wrapping your mind around 4-dimensional data and you’ll understand why dimensionality-reduction is so important. The PCA analysis is just one technique amongst many to do this.
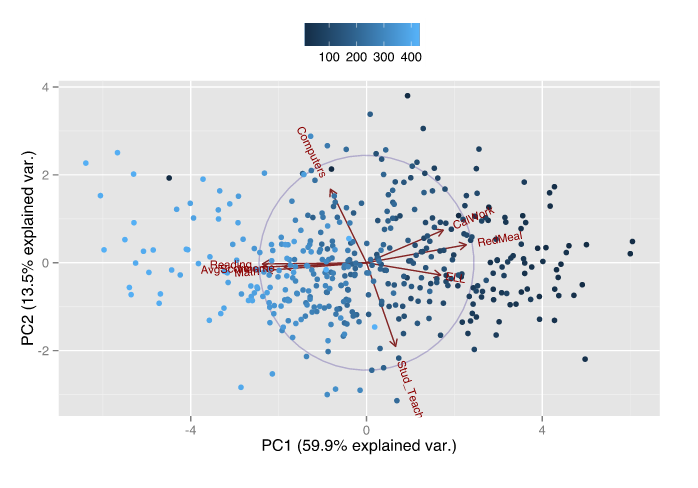
Here is a figure that shows 9-dimensional data projected onto 2 dimensions using PCA. The data is student test scores but also includes socioeconomic data, student:teacher ratio, technology in the classroom, and other indicators. Notice how the lowered socioeconomic indicators (e.g. reduced meal enrollment, English Language Learners, and welfare) are 180º from test performance (Math and Reading scores) showing how socioeconomics and academic performance are highly correlated (and negatively so).
Final Words
While PCA analysis is a quick and simple (relative to other options) technique to reduce the dimensionality of your data–especially when you’re looking for general, preliminary trends, it is far from perfect and should primarily be used to inspect data (not describe it). If there is any reason to assume there might be structure within the data, then a PCA analysis will likely not help you find it. Instead a cluster analysis will be much more fruitful, which you can read my introduction to here. It should also be noted that there exist other dimensionality reducing techniques out there that are based on PCA or use similar methods that may help uncover patterns and trends where PCA fails, but no statistical tool is magic and all hinge on their definition of what their metric for “interesting” is. For example, PCA finds eigenvalues very “interesting”.
I’d love to hear if there is a method that you use or would like to use!
- Can read the wikipedia article on the Characteristic Polynomial of a matrix, or read a more relevant page on eigenpairs.
